New analysis could help uncover potential drug targets for attention deficits and sensory hypersensitivity.
Source: [MIT News Office, Anne Trafton | July 22, 2020]
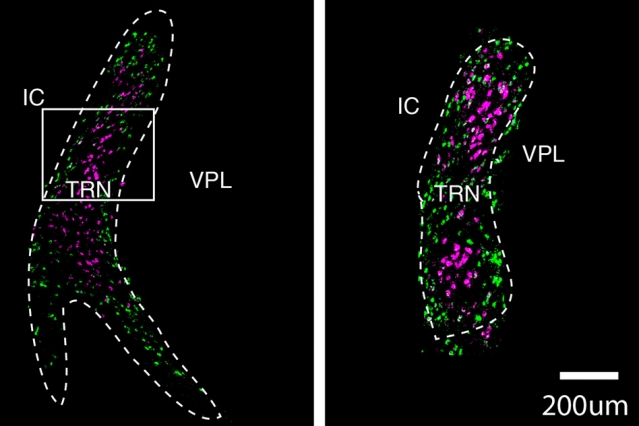
Image: courtesy of the researchers
Many people with autism experience sensory hypersensitivity, attention deficits, and sleep disruption. One brain region that has been implicated in these symptoms is the thalamic reticular nucleus (TRN), which is believed to act as a gatekeeper for sensory information flowing to the cortex.
A team of researchers from MIT and the Broad Institute of MIT and Harvard has now mapped the TRN in unprecedented detail, revealing that the region contains two distinct subnetworks of neurons with different functions. The findings could offer researchers more specific targets for designing drugs that could alleviate some of the sensory, sleep, and attention symptoms of autism, says Guoping Feng, one of the leaders of the research team.
“The idea is that you could very specifically target one group of neurons, without affecting the whole brain and other cognitive functions,” says Feng, the James W. and Patricia Poitras Professor of Neuroscience at MIT and a member of MIT’s McGovern Institute for Brain Research.
Feng; Zhanyan Fu, associate director of neurobiology at the Broad Institute’s Stanley Center for Psychiatric Research; and Joshua Levin, a senior group leader at the Broad Institute, are the senior authors of the study, which appears today in Nature. The paper’s lead authors are former MIT postdoc Yinqing Li, former Broad Institute postdoc Violeta Lopez-Huerta, and Broad Institute research scientist Xian Adiconis.
Distinct populations
When sensory input from the eyes, ears, or other sensory organs arrives in our brains, it goes first to the thalamus, which then relays it to the cortex for higher-level processing. Impairments of these thalamo-cortical circuits can lead to attention deficits, hypersensitivity to noise and other stimuli, and sleep problems.
One of the major pathways that controls information flow between the thalamus and the cortex is the TRN, which is responsible for blocking out distracting sensory input. In 2016, Feng and MIT Assistant Professor Michael Halassa, who is also an author of the new Nature paper, discovered that loss of a gene called Ptchd1 significantly affects TRN function. In boys, loss of this gene, which is carried on the X chromosome, can lead to attention deficits, hyperactivity, aggression, intellectual disability, and autism spectrum disorders.
In that study, the researchers found that when the Ptchd1 gene was knocked out in mice, the animals showed many of the same behavioral defects seen in human patients. When it was knocked out only in the TRN, the mice showed only hyperactivity, attention deficits, and sleep disruption, suggesting that the TRN is responsible for those symptoms.
In the new study, the researchers wanted to try to learn more about the specific types of neurons found in the TRN, in hopes of finding new ways to treat hyperactivity and attention deficits. Currently, those symptoms are most often treated with stimulant drugs such as Ritalin, which have widespread effects throughout the brain.
“Our goal was to find some specific ways to modulate the function of thalamo-cortical output and relate it to neurodevelopmental disorders,” Feng says. “We decided to try using single-cell technology to dissect out what cell types are there, and what genes are expressed. Are there specific genes that are druggable as a target?”
To explore that possibility, the researchers sequenced the messenger RNA molecules found in neurons of the TRN, which reveals genes that are being expressed in those cells. This allowed them to identify hundreds of genes that could be used to differentiate the cells into two subpopulations, based on how strongly they express those particular genes.
They found that one of these cell populations is located in the core of the TRN, while the other forms a very thin layer surrounding the core. These two populations also form connections to different parts of the thalamus, the researchers found. Based on those connections, the researchers hypothesize that cells in the core are involved in relaying sensory information to the brain’s cortex, while cells in the outer layer appear to help coordinate information that comes in through different senses, such as vision and hearing.
“Druggable targets”
The researchers now plan to study the varying roles that these two populations of neurons may have in a variety of neurological symptoms, including attention deficits, hypersensitivity, and sleep disruption. Using genetic and optogenetic techniques, they hope to determine the effects of activating or inhibiting different TRN cell types, or genes expressed in those cells.
“That can help us in the future really develop specific druggable targets that can potentially modulate different functions,” Feng says. “Thalamo-cortical circuits control many different things, such as sensory perception, sleep, attention, and cognition, and it may be that these can be targeted more specifically.”
This approach could also be useful for treating attention or hypersensitivity disorders even when they aren’t caused by defects in TRN function, the researchers say.
“TRN is a target where if you enhance its function, you might be able to correct problems caused by impairments of the thalamo-cortical circuits,” Feng says. “Of course we are far away from the development of any kind of treatment, but the potential is that we can use single-cell technology to not only understand how the brain organizes itself, but also how brain functions can be segregated, allowing you to identify much more specific targets that modulate specific functions.”
The research was funded by the Simons Center for the Social Brain at MIT, the Hock E. Tan and K. Lisa Yang Center for Autism Research at MIT, the James and Patricia Poitras Center for Psychiatric Disorders Research at MIT, the Stanley Center for Psychiatric Research at the Broad Institute, the National Institutes of Health/National Institute for Mental Health, the Klarman Cell Observatory at the Broad Institute, the Pew Foundation, and the Human Frontiers Science Program.
