Tissue-expansion technique could allow scientists to map brain circuits.
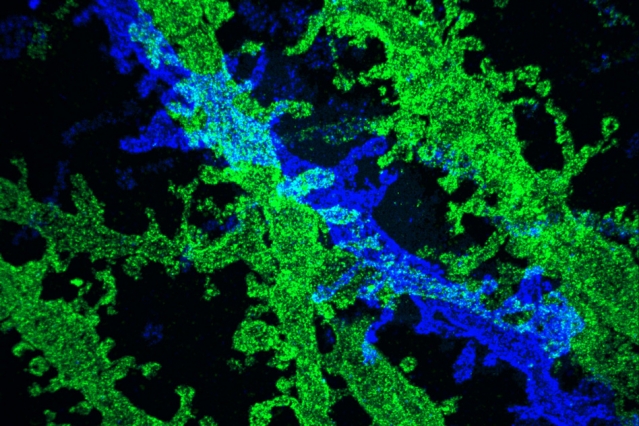
By expanding brain tissue twice, researchers were able to obtain high-resolution images of neurons in the hippocampus.
Image courtesy of the researchers
Source: [Anne Trafton | MIT News Office, April 17, 2017]
The new technique relies on expanding tissue before imaging it with a conventional light microscope. Two years ago, the MIT team showed that it was possible to expand tissue volumes 100-fold, resulting in an image resolution of about 60 nanometers. Now, the researchers have shown that expanding the tissue a second time before imaging can boost the resolution to about 25 nanometers.
This level of resolution allows scientists to see, for example, the proteins that cluster together in complex patterns at brain synapses, helping neurons to communicate with each other. It could also help researchers to map neural circuits, says Ed Boyden, an associate professor of biological engineering and brain and cognitive sciences at MIT.
“We want to be able to trace the wiring of complete brain circuits,” says Boyden, the study’s senior author. “If you could reconstruct a complete brain circuit, maybe you could make a computational model of how it generates complex phenomena like decisions and emotions. Since you can map out the biomolecules that generate electrical pulses within cells and that exchange chemicals between cells, you could potentially model the dynamics of the brain.”
This approach could also be used to image other phenomena such as the interactions between cancer cells and immune cells, to detect pathogens without expensive equipment, and to map the cell types of the body.
Former MIT postdoc Jae-Byum Chang is the first author of the paper, which appears in the April 17 issue of Nature Methods.
Double expansion
To expand tissue samples, the researchers embed them in a dense, evenly generated gel made of polyacrylate, a very absorbent material that’s also used in diapers. Before the gel is formed, the researchers label the cell proteins they want to image, using antibodies that bind to specific targets. These antibodies bear “barcodes” made of DNA, which in turn are attached to cross-linking molecules that bind to the polymers that make up the expandable gel. The researchers then break down the proteins that normally hold the tissue together, allowing the DNA barcodes to expand away from each other as the gel swells.
These enlarged samples can then be labeled with fluorescent probes that bind the DNA barcodes, and imaged with commercially available confocal microscopes, whose resolution is usually limited to hundreds of nanometers.
Using that approach, the researchers were previously able to achieve a resolution of about 60 nanometers. However, “individual biomolecules are much smaller than that, say 5 nanometers or even smaller,” Boyden says. “The original versions of expansion microscopy were useful for many scientific questions but couldn’t equal the performance of the highest-resolution imaging methods such as electron microscopy.”
In their original expansion microscopy study, the researchers found that they could expand the tissue more than 100-fold in volume by reducing the number of cross-linking molecules that hold the polymer in an orderly pattern. However, this made the tissue unstable.
“If you reduce the cross-linker density, the polymers no longer retain their organization during the expansion process,” says Boyden, who is a member of MIT’s Media Lab and McGovern Institute for Brain Research. “You lose the information.”

